Chapter: DNA Replication
Introduction to DNA
DNA is the molecule that stores the hereditary information of life. Every living organism uses DNA as a molecular instruction manual. The information contained within DNA determines the proteins a cell can build, how a cell functions, and ultimately the traits of an organism. In order for life to continue from one cell generation to the next, DNA must be copied with extraordinary accuracy. This process is known as DNA replication.
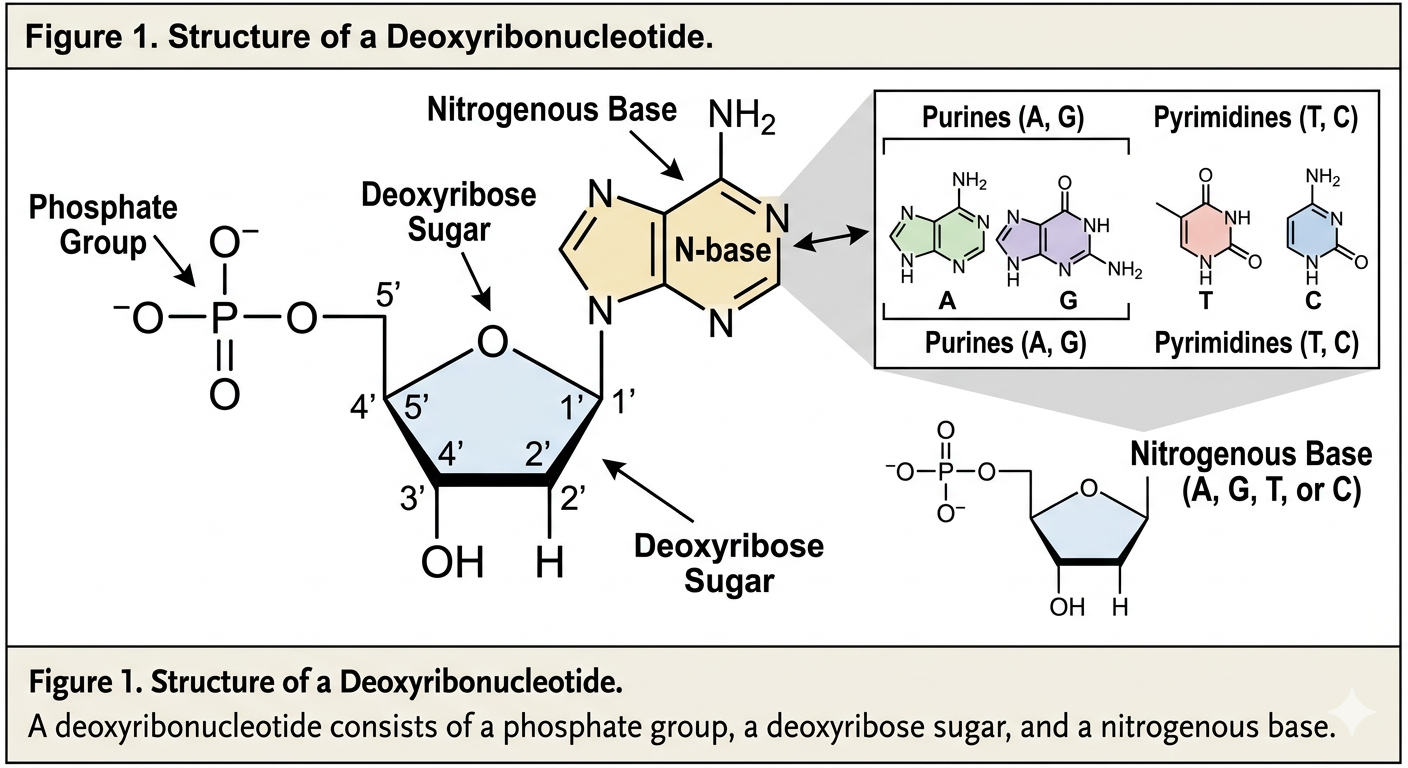
The simplest component (or monomer) of DNA is a nucleotide. More specifically, DNA is composed of deoxyribonucleotides. Each nucleotide consists of three parts: a phosphate group, a five-carbon sugar known as deoxyribose, and a nitrogenous base. The phosphate group is attached to the 5’ carbon of the sugar molecule. Together, alternating sugar and phosphate groups form the structural backbone of DNA. These make up the “sides” of the DNA ladder.
The code of DNA is determined by the nitrogenous bases. There are four nitrogenous bases found in DNA: adenine, thymine, guanine, and cytosine. Adenine and guanine are classified as purines because they contain two carbon-nitrogen rings. Cytosine and thymine are pyrimidines because they contain a single ring.
The DNA Backbone and Directionality
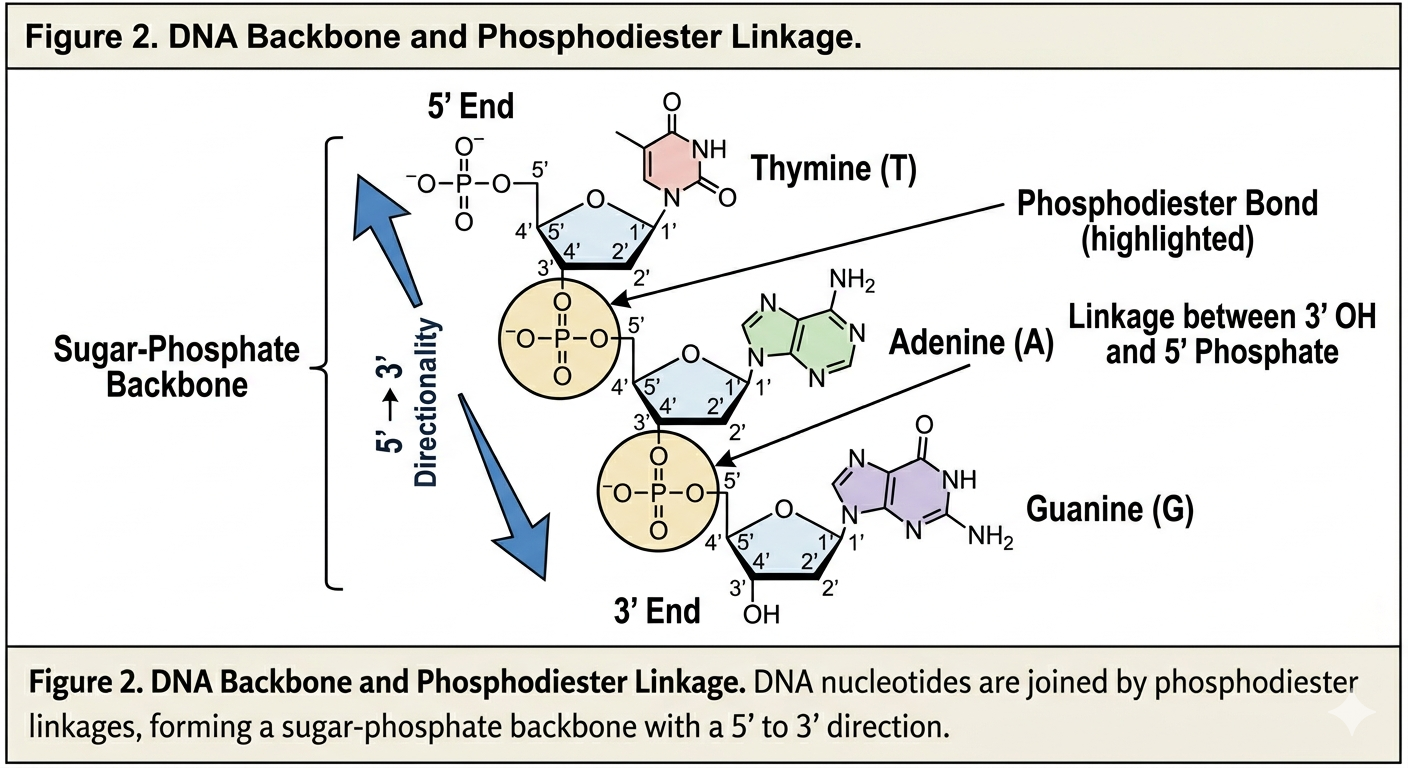
Each nucleotide is connected to the next by a phosphodiester bond. This covalent bond forms between the phosphate group of one nucleotide and the sugar of another nucleotide. Because of this arrangement, DNA has directionality.
One end of a DNA strand contains a free phosphate group attached to the 5’ carbon of the sugar. This is known as the 5’ end. The opposite end contains a free hydroxyl group attached to the 3’ carbon of the sugar. This is known as the 3’ end.
DNA is always synthesized and read in the 5’ to 3’ direction. In many ways, DNA is read like a sentence in a book. We begin at one end and move continuously toward the other. This directionality becomes critically important during DNA replication because enzymes involved in replication can only add nucleotides to the 3’ end of a growing strand.
DNA’s Secondary Structure
Before the 1950s, scientists understood that DNA carried hereditary information, but they did not understand how the molecule was organized. Researchers knew DNA was composed of sugar, phosphate, and nitrogenous bases, but the arrangement of these components remained unclear.
Much of the foundational work concerning DNA structure was performed by James Watson and Francis Crick. In 1953, Watson and Crick proposed the double-helix model of DNA. Their model explained several essential properties of DNA and transformed modern biology.
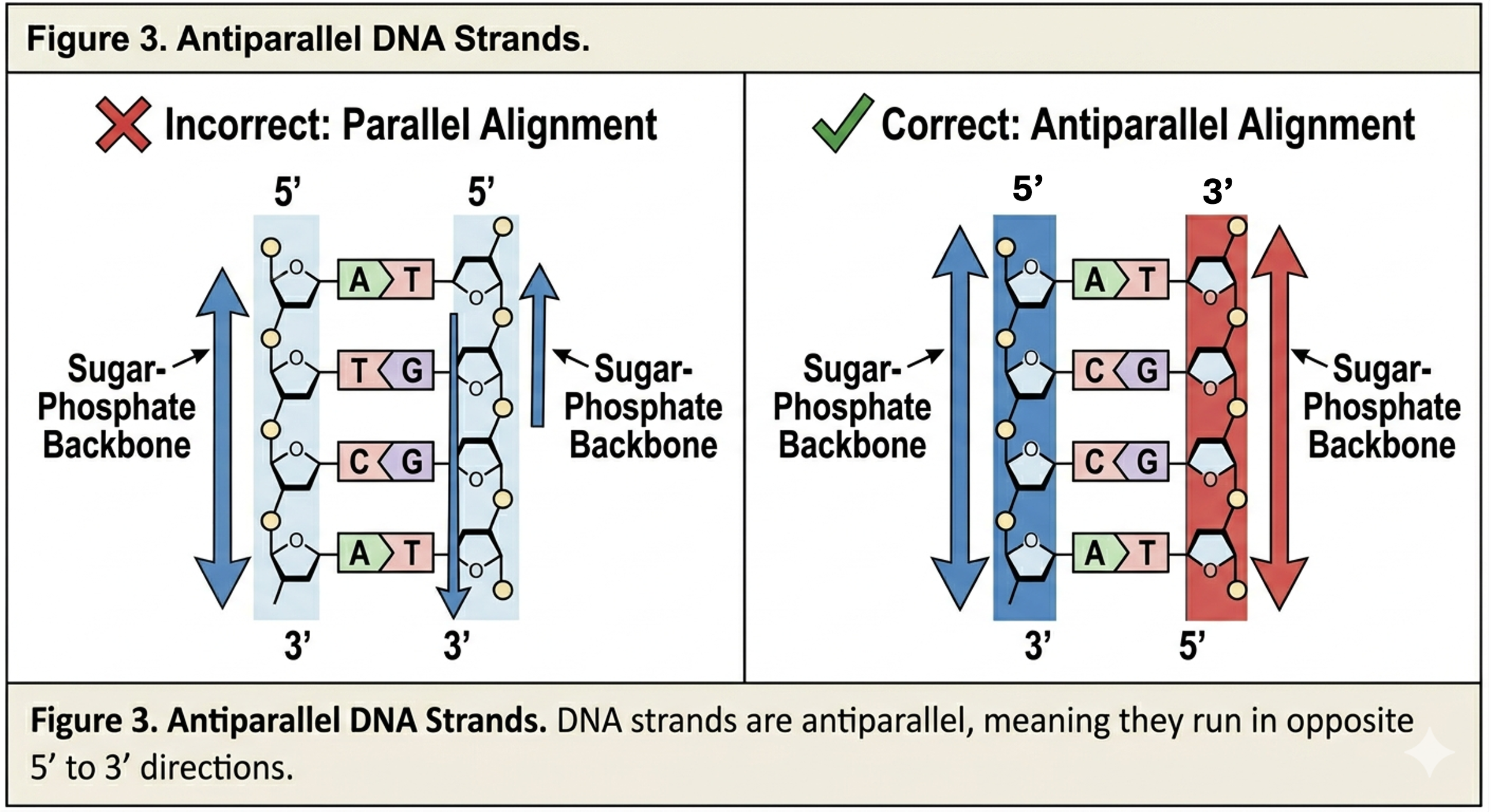
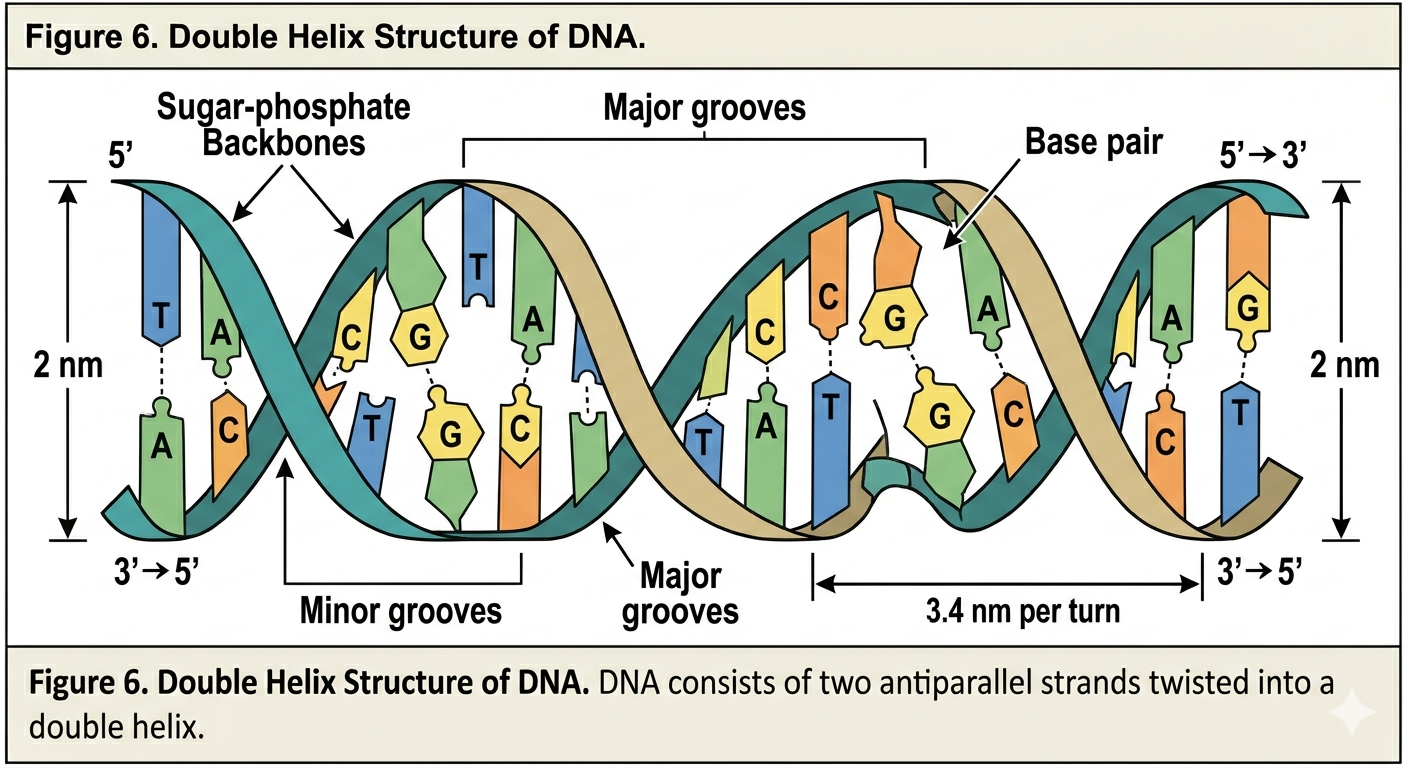
Watson and Crick proposed three major characteristics of DNA structure. First, DNA consists of two strands that twist around each other to form a double helix. Second, the two strands are antiparallel. This means the strands run in opposite directions. One strand runs 5’ to 3’, while the complementary strand runs 3’ to 5’. Antiparallel orientation is essential for replication and base pairing.
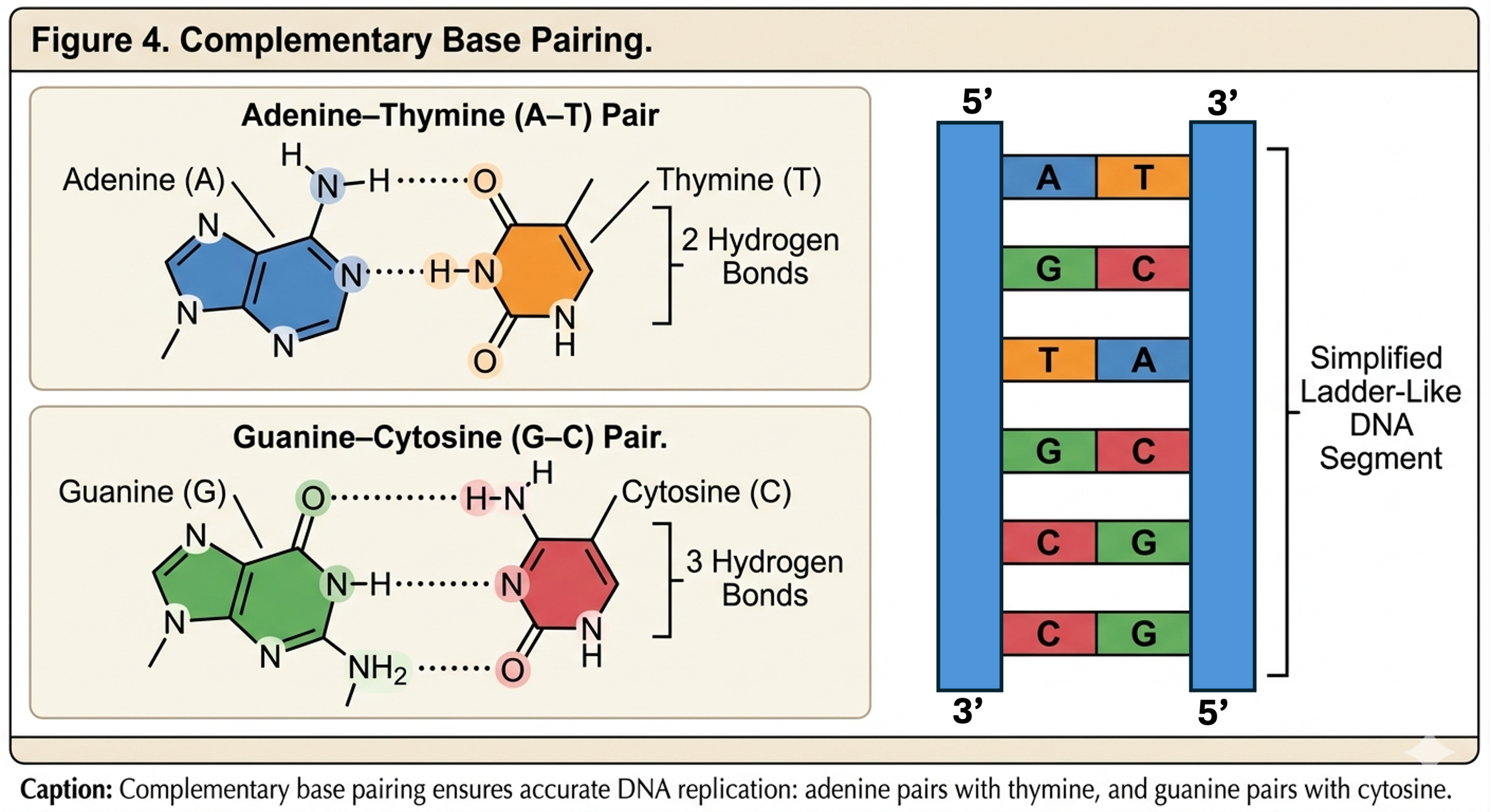
Third, the nitrogenous bases pair according to complementary base pairing rules. Adenine pairs specifically with thymine, while guanine pairs specifically with cytosine. Adenine and thymine are connected by two hydrogen bonds, whereas guanine and cytosine are connected by three hydrogen bonds. The additional bond between guanine and cytosine makes G-C rich regions of DNA more stable.
The complementary nature of DNA immediately suggested a mechanism for copying genetic information. If one strand contains the sequence ATCG, the opposite strand must contain the complementary sequence TAGC. Each strand therefore contains the information necessary to recreate its partner.
Rosalind Franklin and the Discovery of DNA Structure
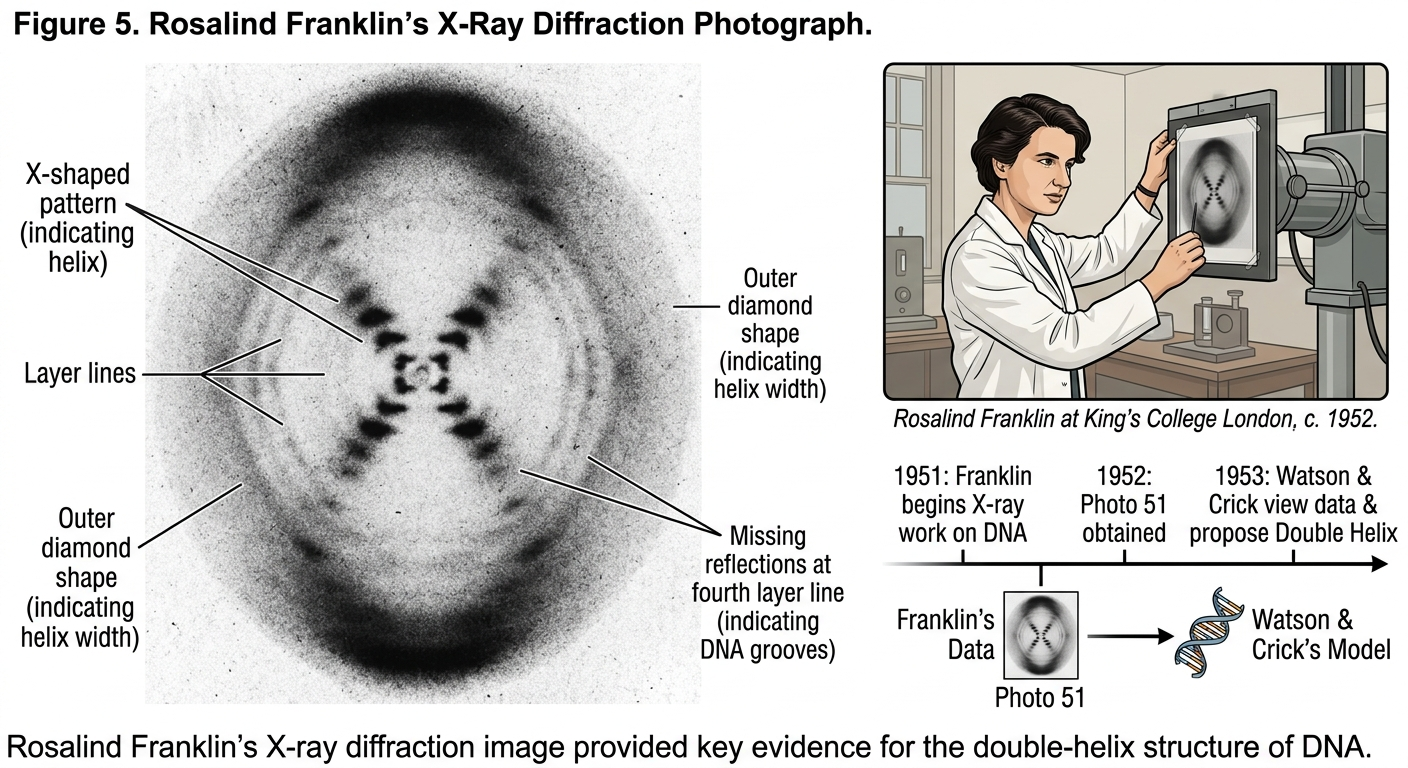
Watson and Crick’s work depended heavily upon the research of chemist Rosalind Franklin. Franklin used X-ray diffraction to study DNA molecules. Her famous image, known as Photograph 51, revealed critical information about DNA’s helical structure.
Franklin’s data demonstrated that DNA had a repeating helical pattern and that the phosphate groups were located on the outside of the molecule. Watson later described seeing Franklin’s image by saying, “My jaw fell open and my pulse began to race.”
Although Watson and Crick received much of the public recognition for discovering DNA structure, Franklin’s experimental work was essential to the breakthrough. Her data were shared without her full permission or knowledge. Franklin was not included as an author on the landmark Nature publication describing the DNA structure. Franklin later developed ovarian cancer and died in 1958 at age 37, likely due in part to radiation exposure from her laboratory research. Because Nobel Prizes are not awarded posthumously, she did not share in the Nobel Prize later awarded to Watson, Crick, and Maurice Wilkins.
DNA as a Template for Replication
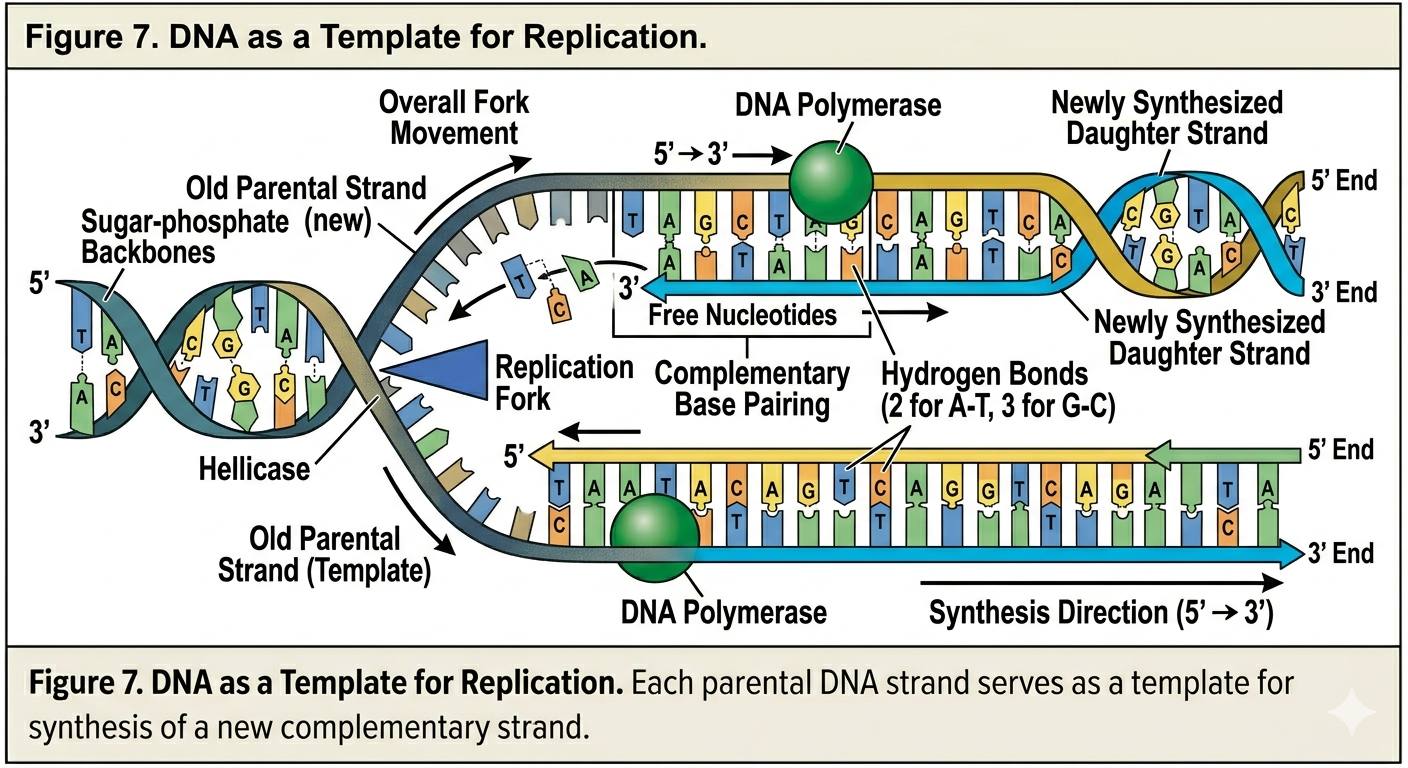
One of the most revolutionary aspects of the Watson and Crick model was the proposal that each DNA strand could serve as a template for the production of a new strand. According to this idea, the two strands of DNA separate from one another during replication. Free nucleotides then pair with exposed bases according to complementary base pairing rules. Adenine pairs with thymine, and guanine pairs with cytosine. In this way, the sequence of one strand determines the sequence of the newly synthesized strand. This mechanism explains how DNA can be copied repeatedly with remarkable accuracy. Every time a cell divides, its DNA must be duplicated so that each daughter cell receives a complete copy of the genome.
Hypotheses of DNA Replication
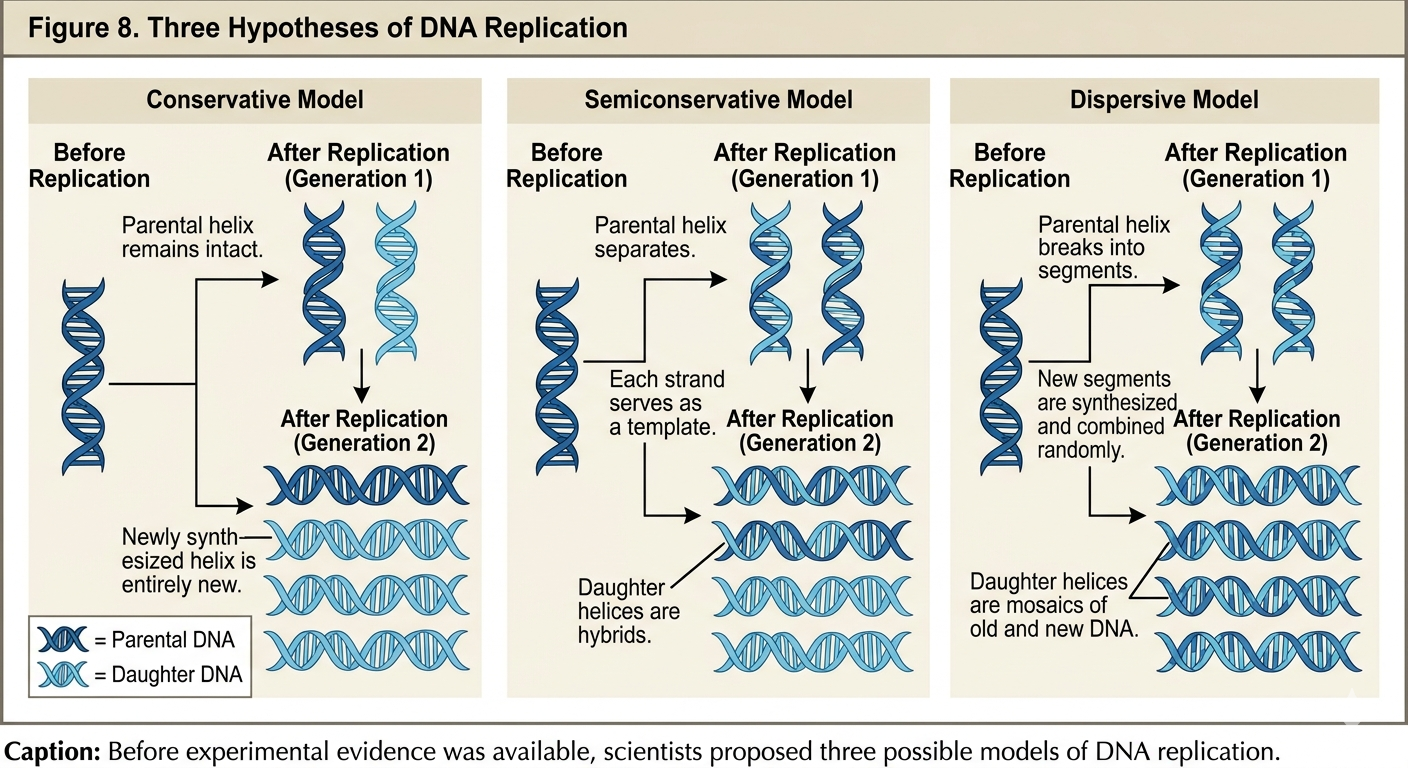
Once the structure of DNA had been established, scientists next attempted to determine precisely how DNA replication occurred. Three competing hypotheses were proposed.
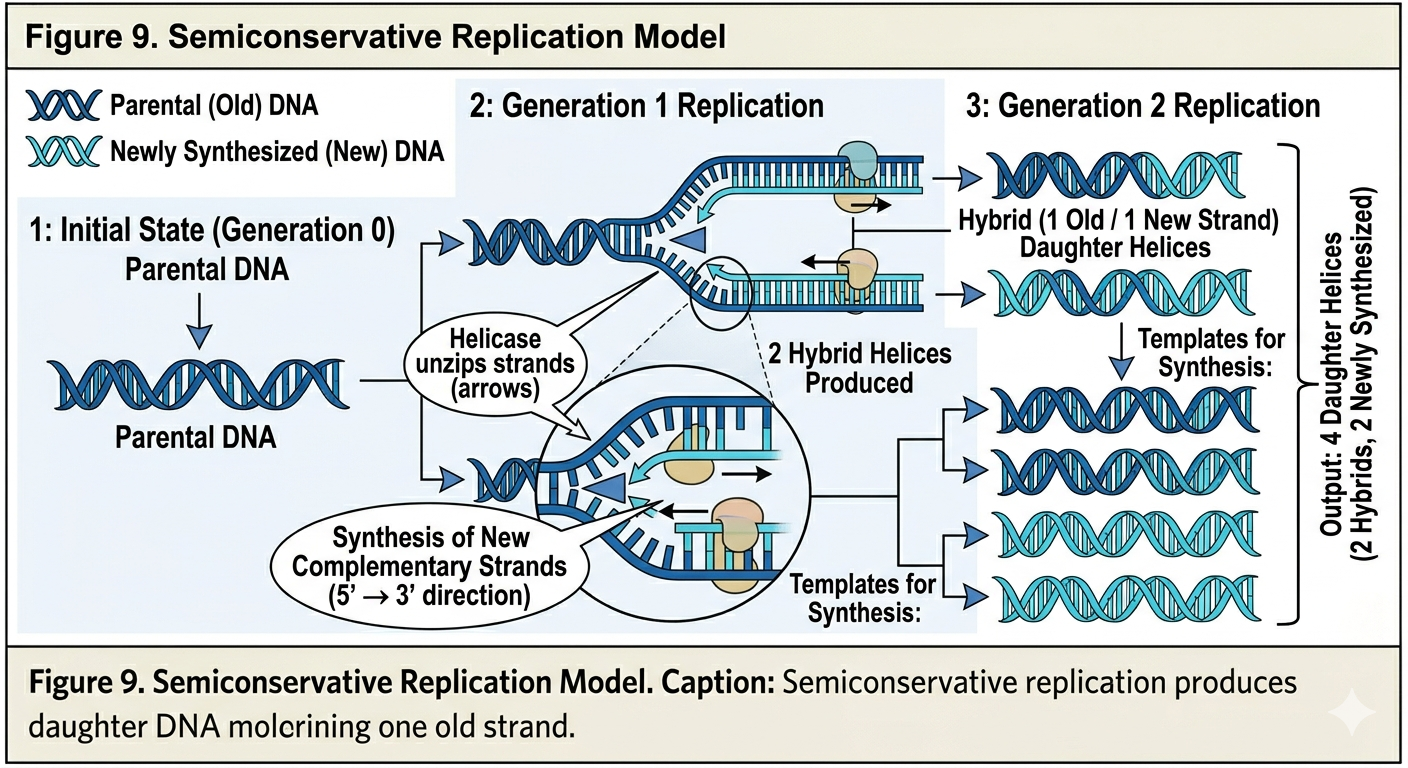
Semiconservative Replication
The semiconservative hypothesis proposed that the two parental DNA strands separate from one another. Each original strand then serves as a template for the synthesis of a new complementary strand. After replication, each daughter DNA molecule contains one original parental strand and one newly synthesized strand. Thus, half of the original molecule is conserved in each new DNA double helix.
Conservative Replication
The conservative hypothesis proposed that the parental DNA double helix remains completely intact during replication. According to this model, the parental DNA molecule would serve as a template for an entirely new double-stranded DNA molecule. After replication, one DNA molecule would consist entirely of the original parental DNA, while the second molecule would consist entirely of newly synthesized DNA.
Dispersive Replication
The dispersive hypothesis proposed that parental DNA molecules were fragmented during replication. According to this model, each strand of daughter DNA would contain alternating segments of old parental DNA and newly synthesized DNA. Rather than preserving entire parental strands, the original DNA would become dispersed throughout the newly synthesized molecules.
The Meselson–Stahl Experiment
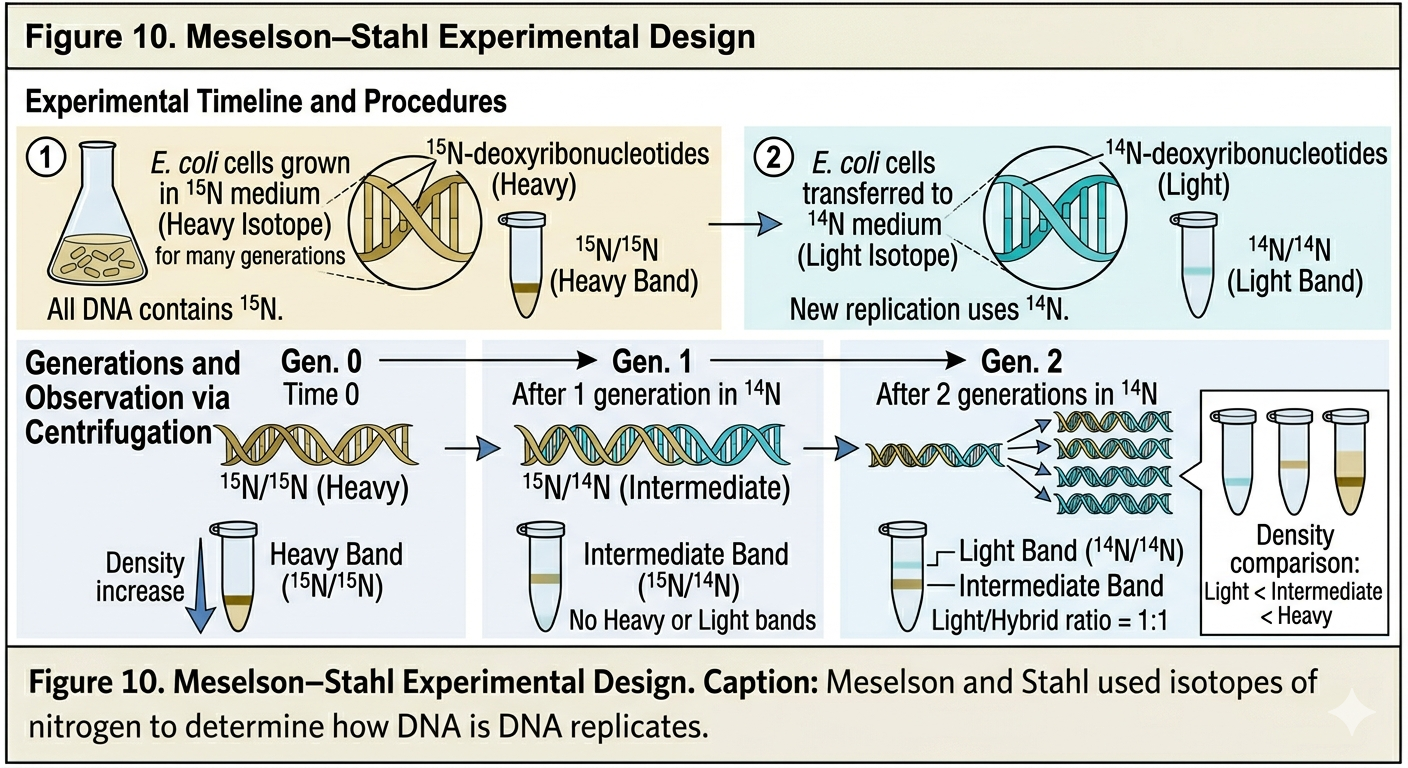
In 1958, Matthew Meselson and Franklin Stahl designed one of the most elegant experiments in biology to determine which replication hypothesis was correct. Nitrogen is a major component of DNA bases. Most naturally occurring nitrogen exists as the isotope 14N, but a heavier stable isotope, 15N, can also be incorporated into DNA. Meselson and Stahl grew E. coli bacteria for many generations in a medium containing only 15N. As a result, the bacterial DNA became labeled entirely with heavy nitrogen. The bacteria were then transferred to a medium containing normal 14N. As the bacteria reproduced, newly synthesized DNA strands incorporated 14N rather than 15N. The researchers extracted DNA after successive generations and separated the DNA using density-gradient centrifugation. DNA containing 15N is denser than DNA containing 14N, allowing the researchers to determine the composition of the DNA molecules.
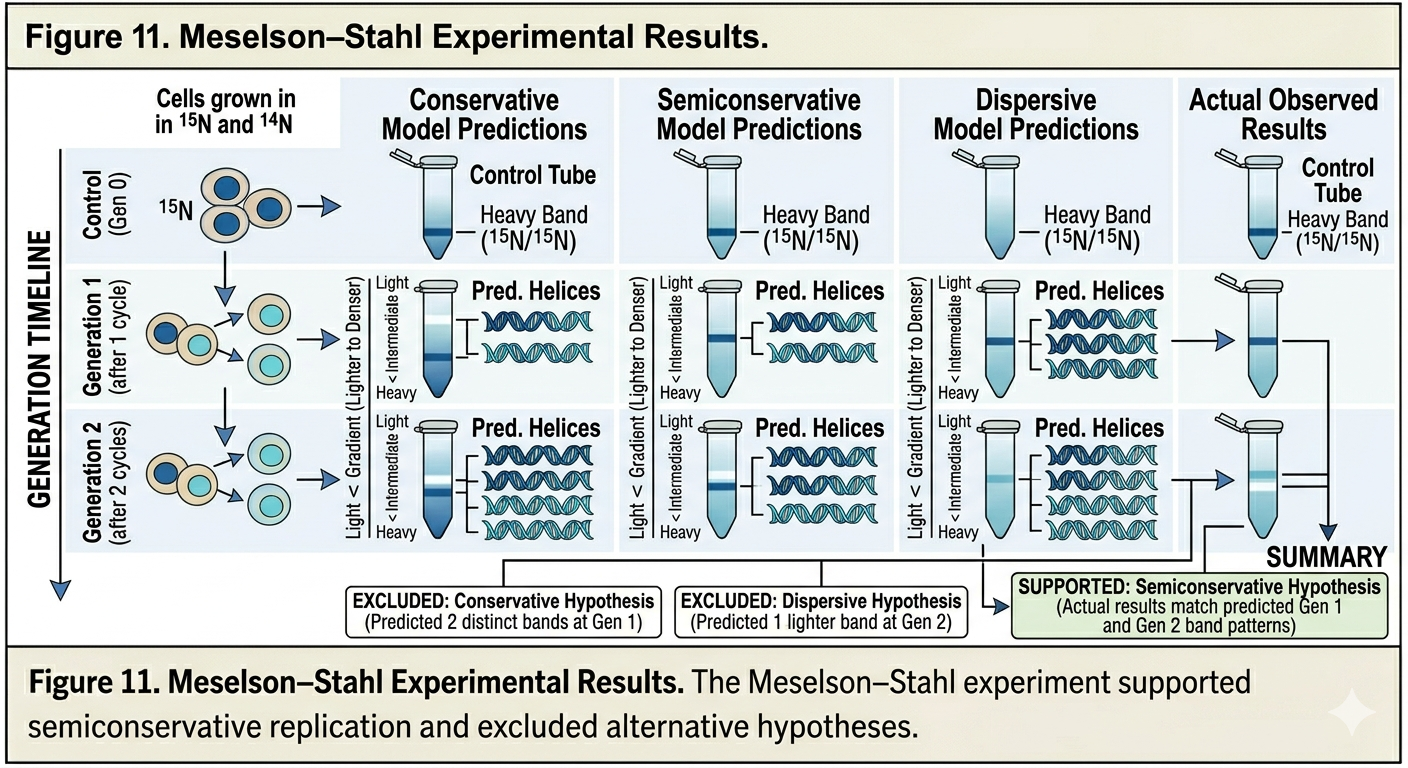
Predictions of the Three Models
Each replication hypothesis predicted different results. After one generation, the conservative model predicted two bands would appear: one heavy band containing entirely parental DNA and one light band containing entirely new DNA. The semiconservative model predicted that after one generation, all DNA would possess intermediate density because each molecule would contain one 15N strand and one 14N strand. The dispersive model also predicted intermediate-density DNA after one generation because each strand would contain mixtures of 15N and 14N. After two generations, however, the predictions differed. The semiconservative model predicted two bands: one intermediate-density band and one light-density band. The dispersive model predicted only a single band that gradually became lighter over successive generations.
Meselson-Stahl Experimental Results
After one generation in 14N medium, Meselson and Stahl observed a single intermediate-density band. This immediately ruled out conservative replication. After two generations, two bands appeared: one intermediate-density band and one light-density band. This result was consistent only with semiconservative replication. The experiment demonstrated conclusively that DNA replication is semiconservative. Each daughter DNA molecule contains one original parental strand and one newly synthesized strand. The Meselson–Stahl experiment remains one of the most important experiments in molecular biology because it provided direct evidence for the mechanism of DNA replication.
DNA Polymerase
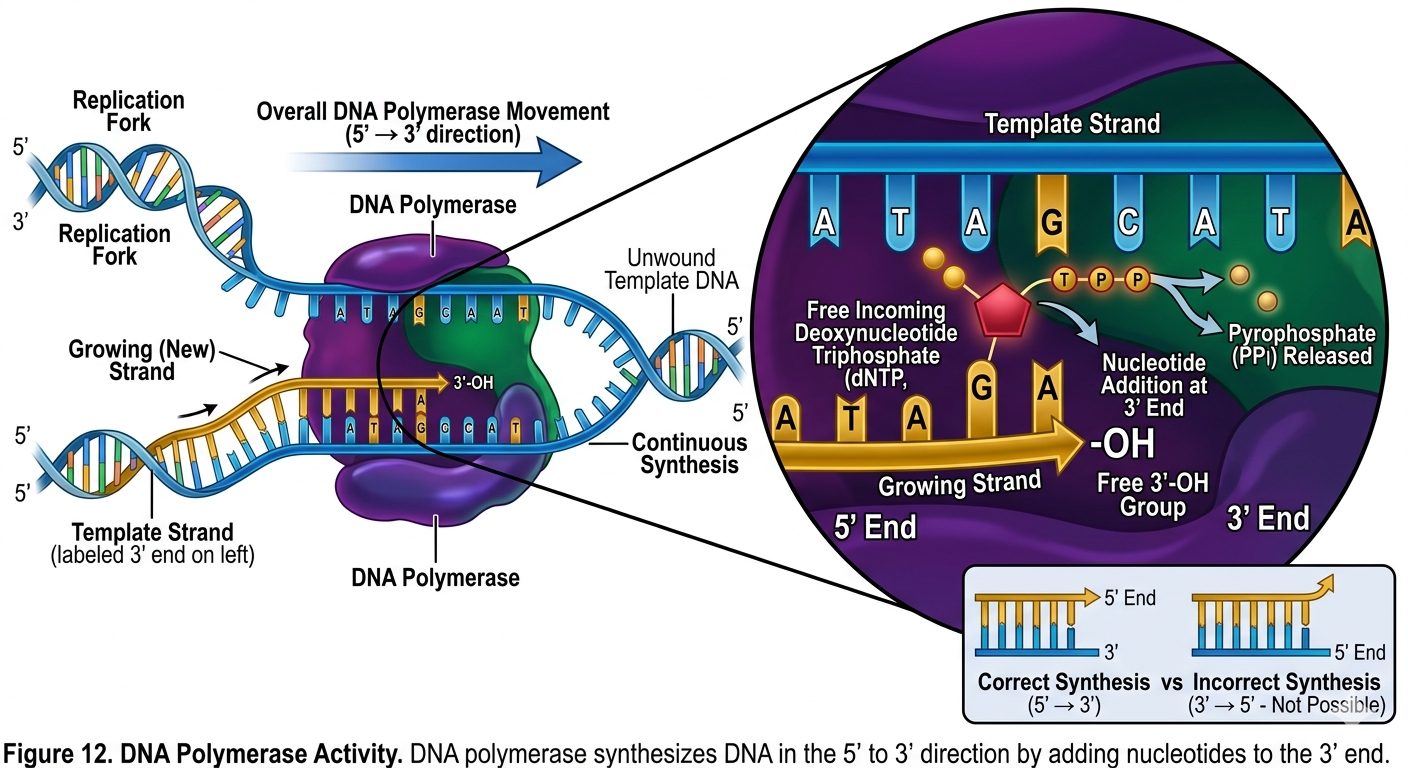
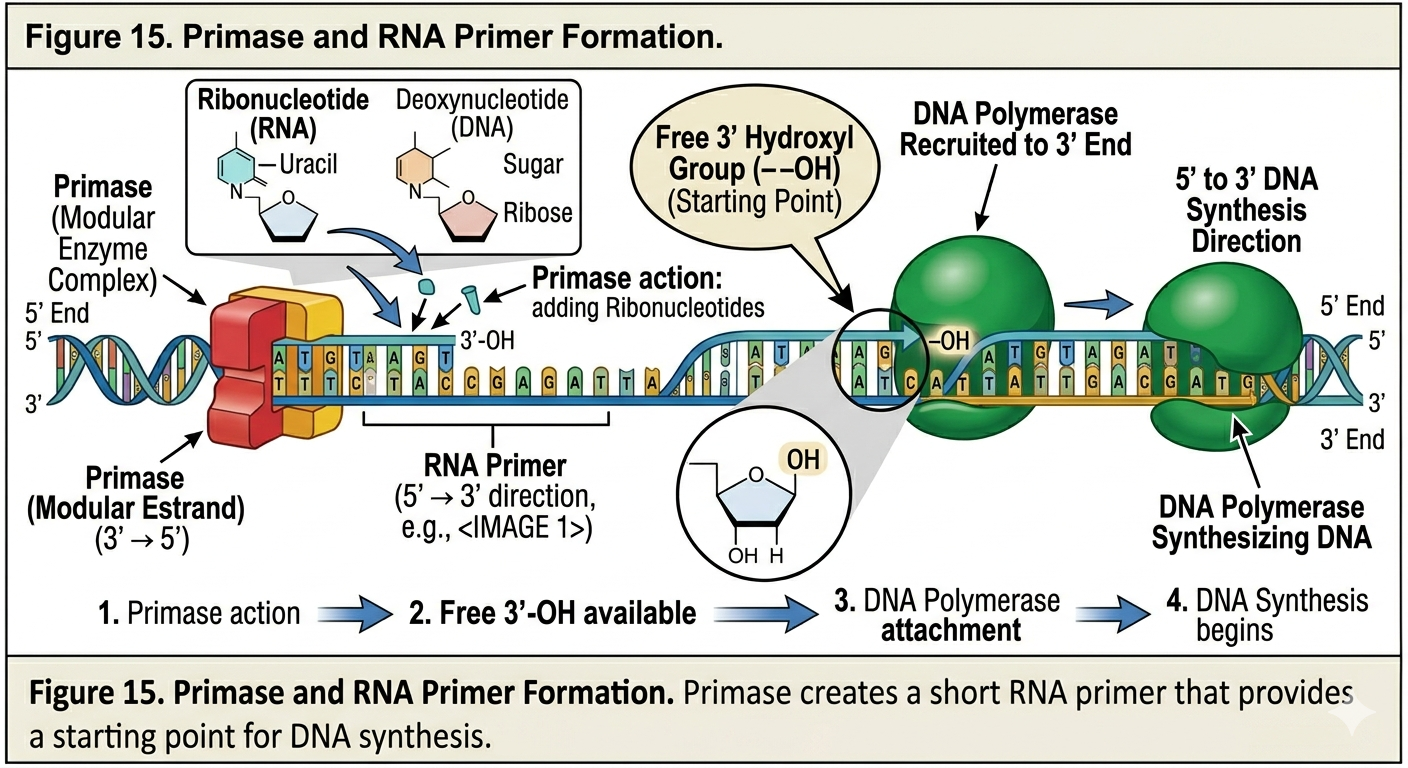
The discovery of DNA polymerase provided the molecular mechanism explaining how DNA replication occurs. DNA polymerases are enzymes that synthesize new DNA strands by adding nucleotides to an existing strand. DNA polymerase can only add nucleotides to the 3’ hydroxyl end of a growing DNA strand. Consequently, all DNA synthesis occurs in the 5’ to 3’ direction. DNA polymerase does not begin synthesis from scratch. It requires an existing nucleotide chain with a free 3’ hydroxyl group. This requirement explains the need for RNA primers during DNA replication, more on that in a minute.
Different organisms possess several types of DNA polymerases. In bacteria, DNA polymerase III performs most DNA synthesis during replication, while DNA polymerase I removes RNA primers and replaces them with DNA. In eukaryotes, multiple specialized DNA polymerases participate in replication and repair.
Origins of Replication and Replication Bubbles
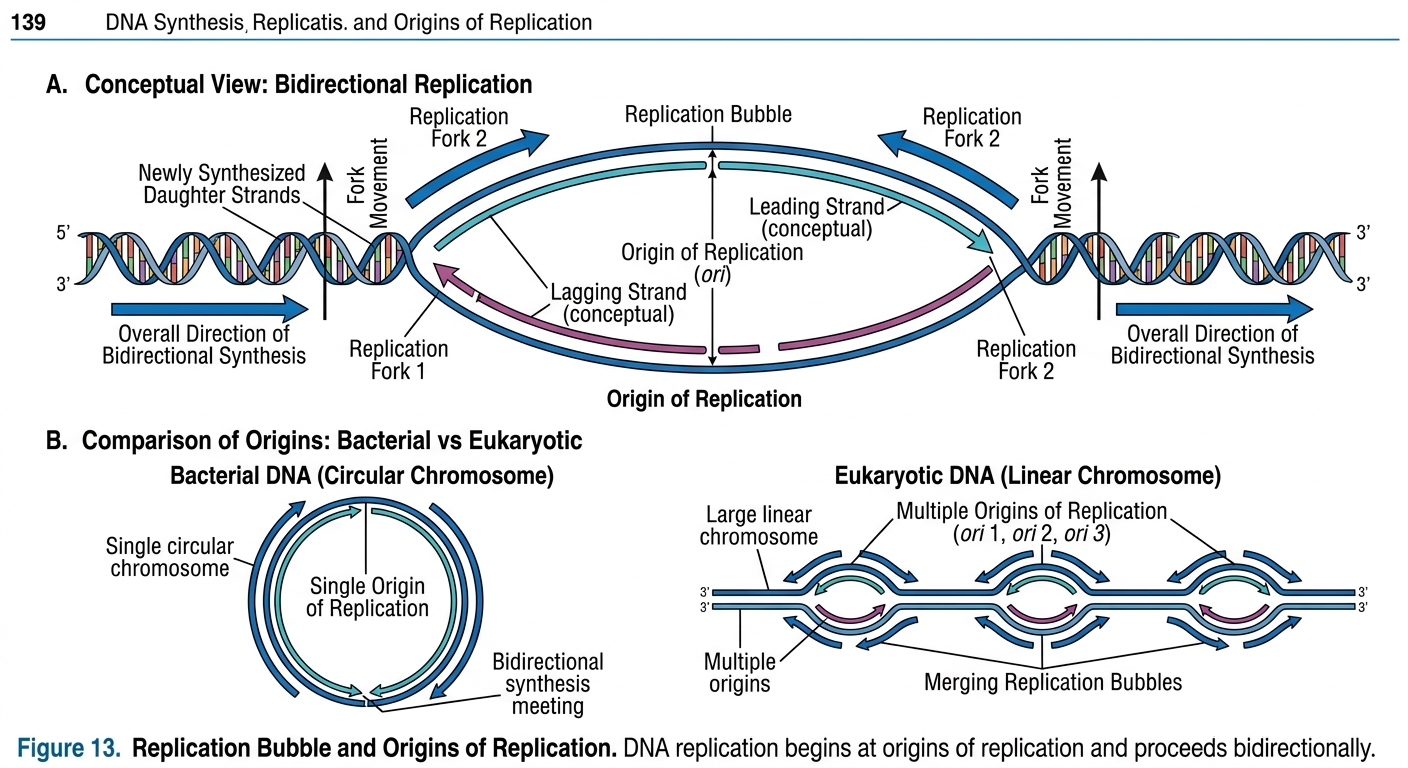
DNA replication begins at specific sites called origins of replication. At these locations, proteins bind to DNA and initiate the opening of the double helix.
Bacteria
In bacteria, which possess circular DNA, replication typically begins at a single origin of replication.
Eukaryotes
Eukaryotic DNA is linear, known as chromosomes. Because of their enormous size, replication begins simultaneously at many origins along each chromosome. This allows the entire genome to be copied efficiently within a reasonable amount of time. As DNA opens at an origin, a replication bubble forms. Replication proceeds outward in both directions from the origin, producing two replication forks.
Initiation of DNA Replication
Let’s look at the details of how DNA is replicated. DNA begins replication by being opened, unwound and primed for replication.
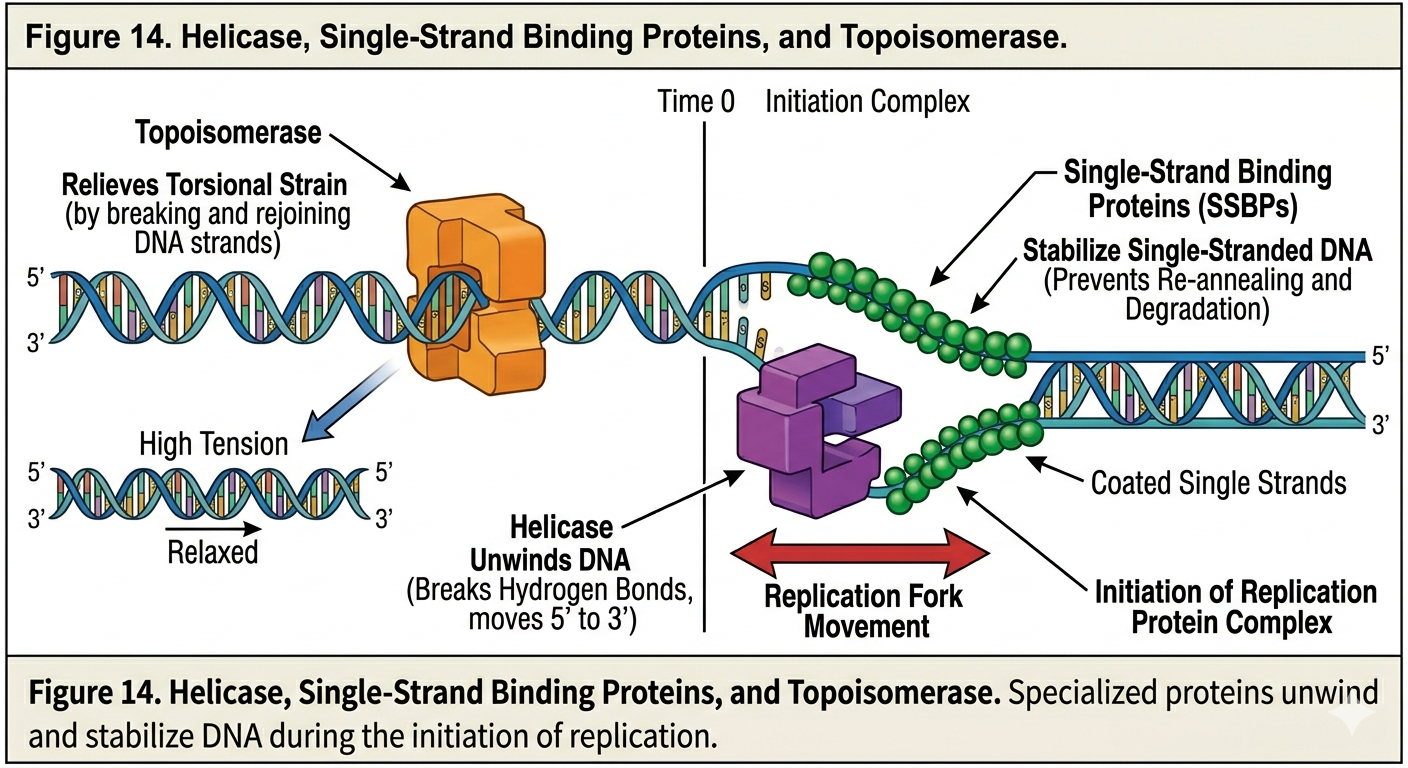
Helicase
DNA replication begins by opening and preparing the double helix. The enzyme helicase unwinds the DNA molecule by breaking the hydrogen bonds between complementary bases. This separates the two DNA strands and creates replication forks.
Single Stranded Binding Proteins
Once the strands separate, proteins known as single-strand binding proteins attach to the exposed DNA strands. These proteins stabilize the separated strands and prevent them from reattaching via complementary base pairing.
Topoisomerase
As DNA unwinds, tension builds ahead of the replication fork. The twisting strain resembles attempting to unwind a tightly coiled telephone cord or slinky. To relieve this tension, enzymes known as topoisomerases cut, unwind, and rejoin DNA ahead of the replication fork. Without topoisomerase activity, DNA would become excessively supercoiled and replication would eventually stop.
Primase
DNA polymerase cannot initiate synthesis independently. A specialized enzyme called primase synthesizes short sequence of RNA, known as RNA primers complementary to the DNA template. These primers provide the free 3’ hydroxyl group required for DNA polymerase to begin replication. Once primers are in place, DNA replication can proceed.
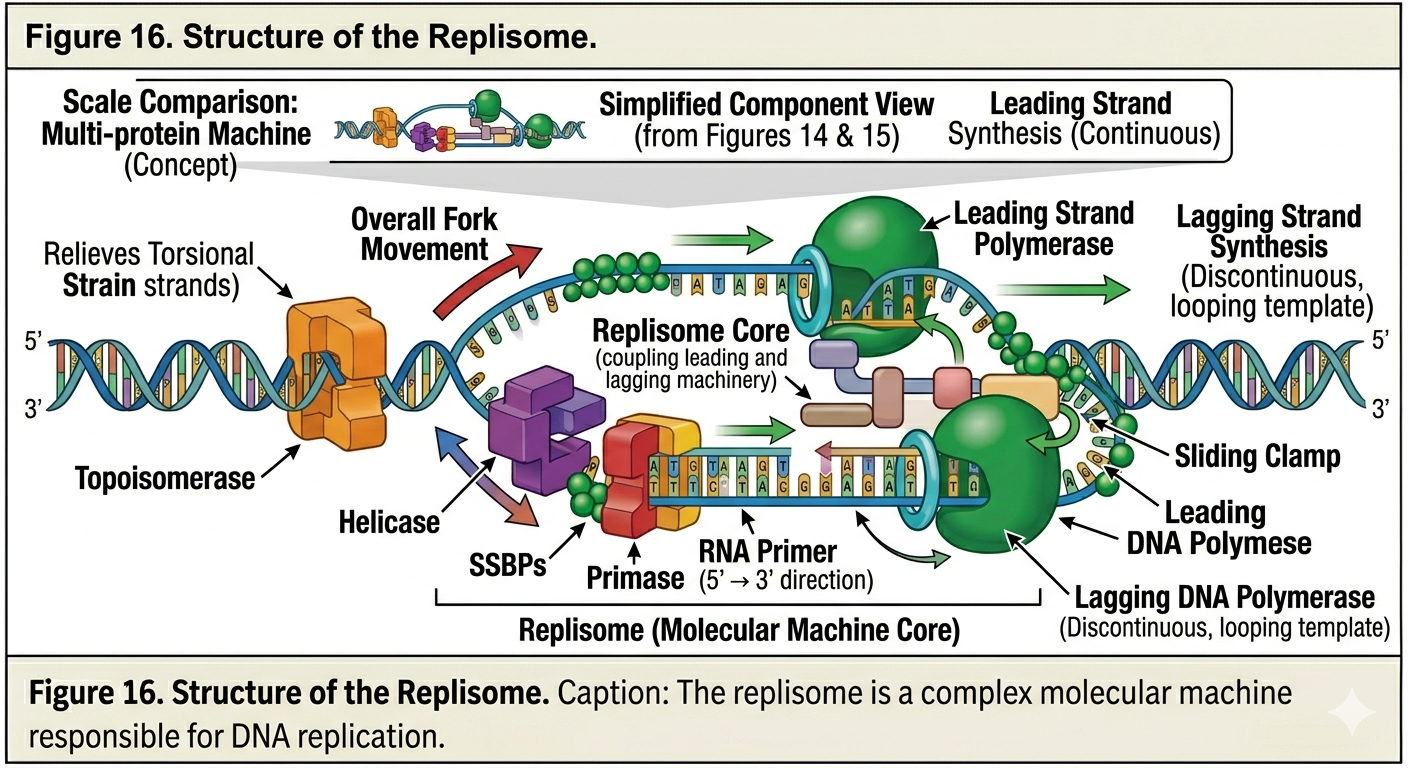
The Replisome
DNA replication is carried out by a massive protein complex known as the replisome. The replisome coordinates the activities of helicase, primase, DNA polymerase, and other associated proteins. Because DNA strands are antiparallel and DNA polymerase synthesizes only in the 5’ to 3’ direction, the two strands are copied differently.
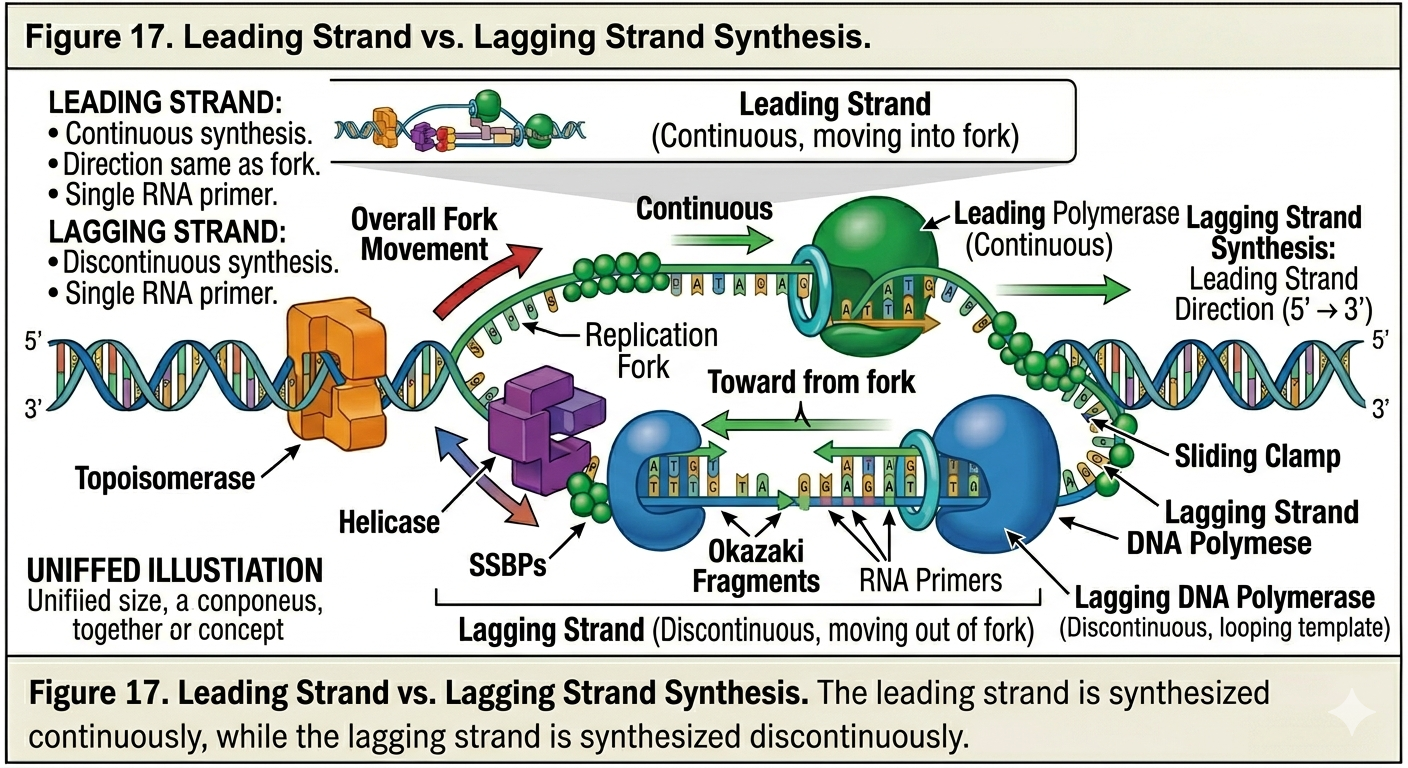
Leading Strand Synthesis
The leading strand is synthesized continuously toward the replication fork. Because the template strand runs in the 3’ to 5’ direction relative to fork movement, DNA polymerase can continuously add nucleotides as the fork opens. Only a single RNA primer is typically required for synthesis of the leading strand.
Lagging Strand Synthesis
The lagging strand is synthesized discontinuously away from the replication fork. Because DNA polymerase can synthesize DNA only in the 5’ to 3’ direction, the lagging strand must be copied in short segments as more template DNA becomes exposed. Primase repeatedly synthesizes new RNA primers as the replication fork opens. DNA polymerase extends these primers to form short DNA fragments known as Okazaki fragments. Each fragment is synthesized away from the replication fork. The lagging strand therefore requires repeated cycles of priming and synthesis.
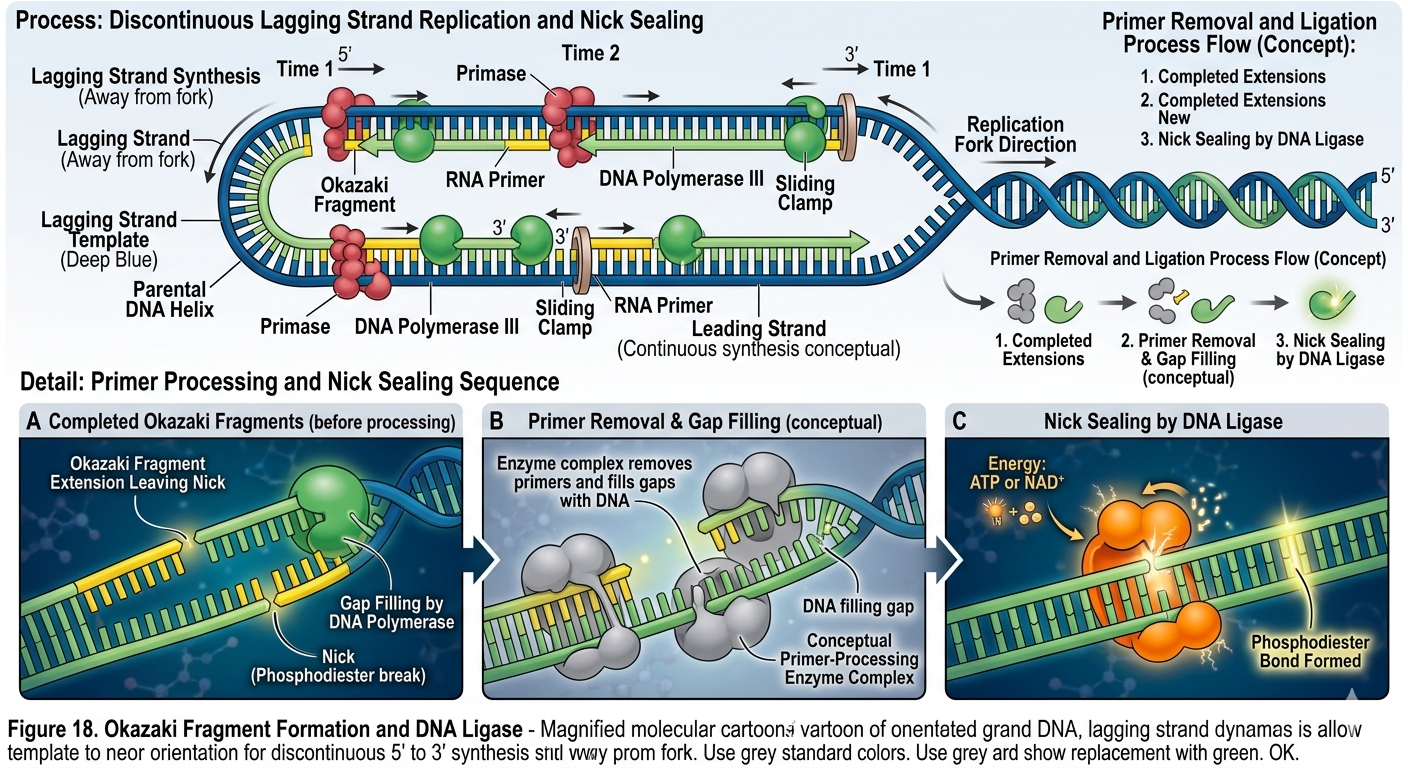
Processing Okazaki Fragments
Once Okazaki fragments are synthesized, the RNA primers must be removed and replaced with DNA. In bacteria, DNA polymerase I removes RNA primers and fills the resulting gaps with DNA nucleotides. The remaining breaks between fragments are sealed by DNA ligase. DNA ligase forms phosphodiester bonds between adjacent DNA fragments, creating a continuous sugar-phosphate backbone. Without DNA ligase, the lagging strand would remain fragmented.
Replication in Prokaryotes and Eukaryotes
Although the basic mechanism of DNA replication is highly conserved, several important differences exist between prokaryotic and eukaryotic replication. Prokaryotic DNA is circular and possess a single origin of replication. Replication proceeds rapidly because bacterial genomes are relatively small. Eukaryotic chromosomes are linear and much larger. Multiple origins of replication are necessary to complete DNA replication efficiently. Eukaryotic DNA is also tightly associated with histone proteins to form chromatin. During replication, nucleosomes must be temporarily disassembled and then reassembled onto newly synthesized DNA. Additionally, eukaryotic cells face a unique challenge associated with replicating the ends of linear chromosomes.
Telomeres and Telomerase
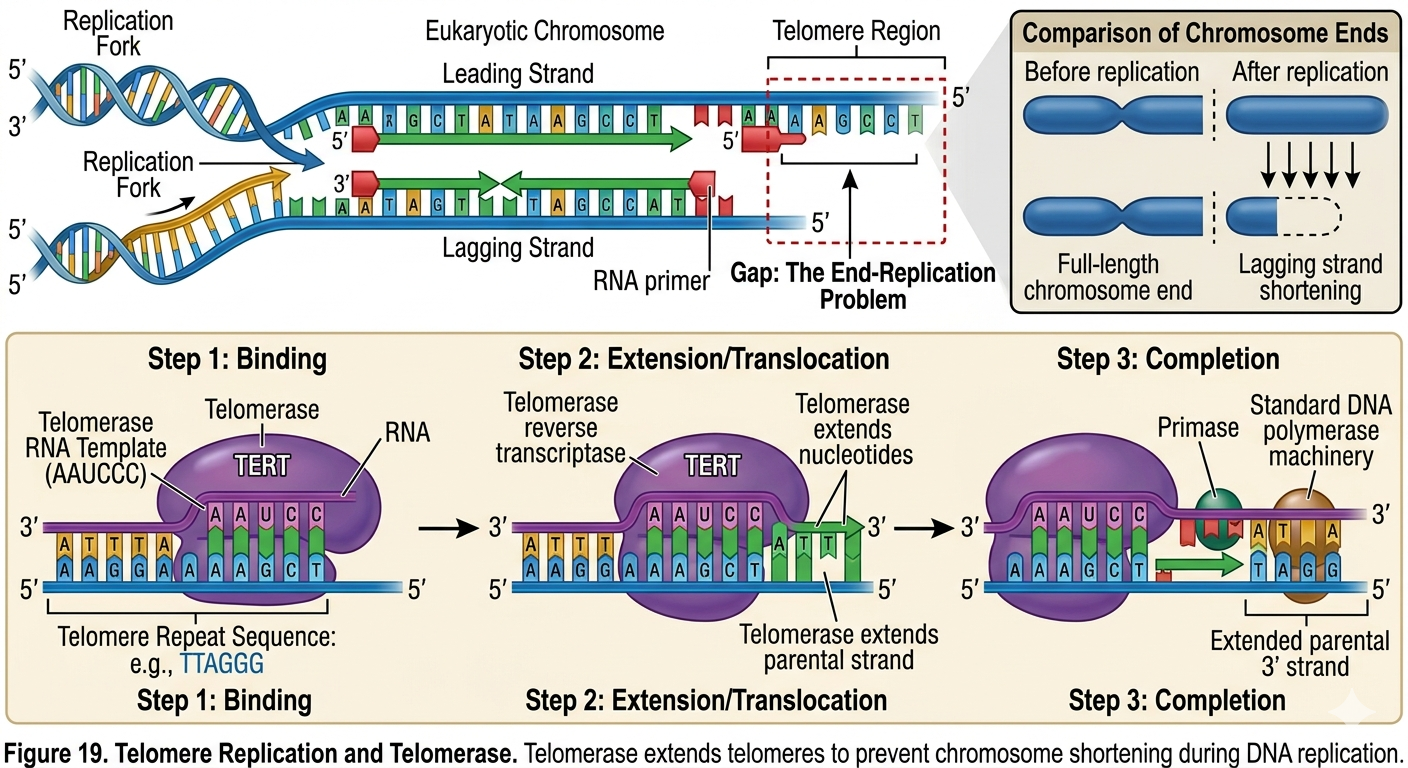
The ends of eukaryotic chromosomes are known as telomeres. Telomeres contain repetitive noncoding DNA sequences that protect chromosomes from degradation. DNA replication creates a problem at the ends of linear chromosomes. Once the final RNA primer on the lagging strand is removed, DNA polymerase cannot replace it because there is no available upstream 3’ hydroxyl group. As a result, chromosomes would become slightly shorter during every round of replication. To prevent the loss of important genetic information, cells use an enzyme called telomerase.
Telomerase contains its own RNA template and extends telomeres by adding repetitive DNA sequences to chromosome ends. This extension allows additional Okazaki fragments to be synthesized, preventing excessive chromosome shortening. In most somatic cells, telomerase activity is low or absent. Consequently, telomeres gradually shorten with age and repeated cell division. In contrast, stem cells, germ cells, and many cancer cells possess active telomerase, allowing them to divide repeatedly.
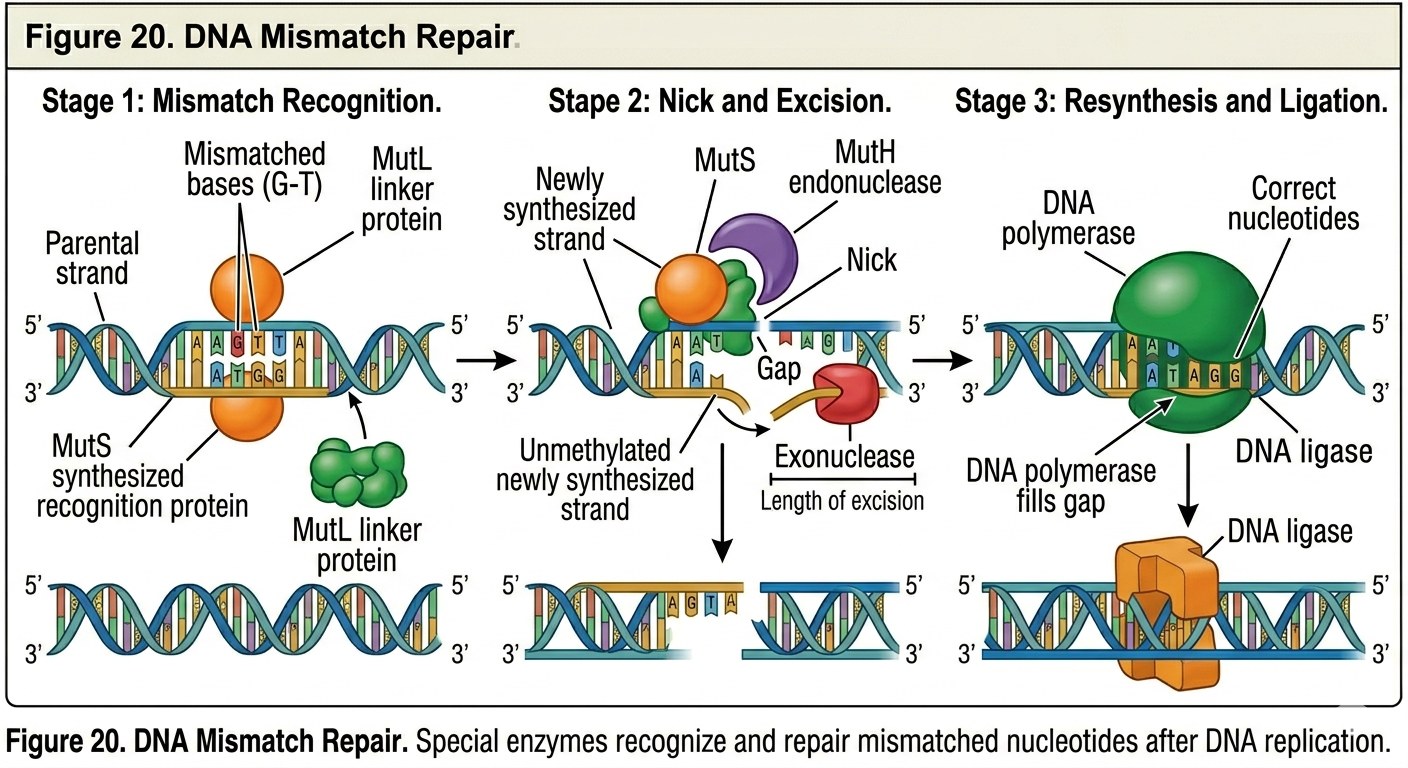
Proofreading and DNA Repair
DNA replication is extraordinarily accurate. Only approximately one mistake occurs for every billion nucleotides copied. This accuracy results from several mechanisms. First, complementary base pairing itself is highly selective. Adenine pairs most efficiently with thymine, and guanine pairs most efficiently with cytosine. Second, many DNA polymerases possess proofreading ability. As DNA polymerase synthesizes DNA, it continuously checks newly added nucleotides. If an incorrect nucleotide is inserted, the enzyme removes it and replaces it with the correct base. These repair systems help maintain genome stability and reduce mutation rates.
Mutations
Despite the remarkable accuracy of DNA replication and repair, errors still occasionally occur. Permanent changes in DNA sequence are known as mutations. Some mutations are silent and produce no detectable effect. Others may alter protein function and cause disease. Certain mutations can provide evolutionary advantages. Over long periods of time, mutations serve as the raw material for evolution by introducing genetic variation into populations. Although mutations are often harmful or neutral, natural selection occasionally favors beneficial mutations that improve survival or reproduction.
The Importance of DNA Replication
DNA replication is one of the most essential processes in biology. Without accurate replication, cells could not divide, organisms could not grow, and hereditary information could not be passed from one generation to the next. The semiconservative mechanism of DNA replication ensures that genetic information is copied faithfully while still allowing the possibility of variation through mutation. The coordinated action of helicase, primase, DNA polymerase, ligase, topoisomerase, and repair enzymes allows cells to replicate billions of nucleotides with extraordinary precision.
Modern genetics, biotechnology, molecular medicine, and evolutionary biology all depend upon our understanding of DNA replication. From cancer research to forensic science to genetic engineering, the study of DNA replication remains central to modern biology.